At this time, we have the program incorporated into usmapmaker.pl as a "resolution enhancement option". This option performs downscaling of PRISM-derived degree-day maps for the 5-state Northwestern US. The downscaling now takes 1:15 (1 minute 15 second or about 1.5 KM) heat unit data down to 0:09 (9 second) resolution using USGS GTOPO30 DEM (30 second elevation, about 580 meter resolution) and 360m DEM. This is approximately a 16 fold improved resolution, and does improve both relative and absolute precision and accuracy, as well as the readability, of the degree-day maps. The method uses a moving window 5x5 weighted simple linear regression. In addition, it has a) ability to ignore data beyond map edges, and b) a threshold r-value with "fail-back" to the non-downscaled value (currently used at r=0.15). Both distance-weights and threshold r-values are user-determined and set.
This program may provide a tool for other downscaling needs to provide an alternative to gaussian smoothing.
A brief outline of the process for computing PRISM-derived degree-day maps without the downscaling step is available from /wea/avgmapexp.html, and documentation for using the mapmaker application is available from /wea/mapmkrdoc.html. Please review.
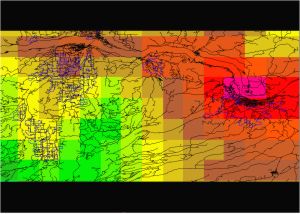
Fig. 1. Initial PRISM-derived DD map, 4 km (corrected using near-realtime site data) |
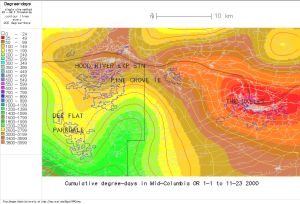 Fig. 2. Final (gaussian-smoothed) DD map, 500 m (as available from usmapmaker.pl web application with the default, "original resolution" option selected) |
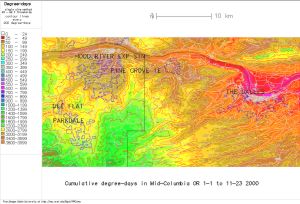 Fig. 3. New downscaled DD map, 100 m (downscaled using r.downscale.pl, no additional gaussian smoothing, this based on USGS 30m DEM, no longer available at mapmaker.pl) |
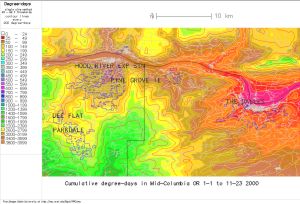 Fig. 4. New downscaled DD map, 500 m (downscaled using r.downscale.pl, with additional gaussian smoothing, based on USGS GTOPO30 DEM, now available at mapmaker.pl as the "resolution enhancement" option) |
r.downscale.pl: Perl script to use r.mapcalc and GWR to downscale from low to high
resolution using 5 x 5 local weighted regressions between variables
like rainfall or temperature (dep. variable) and elevation
(indep. variable).
The program expects to be run twice:
Once at the lower resolution to build models using calc=BUILD,
then at a higher resolution using calc=EST to calculate new
high resolution values. Example usage:
GRASS >g.region res=0:02:30.00 # set lower res
GRASS >r.downscale.pl input=H_50 elev_lo=lodem calc=BUILD # build model
GRASS >g.region res=0:00:09.00 # set higher res
GRASS >r.downscale.pl elev_hi=DEM30m elev_lo=lodem input=H_50 output=HI_50 calc=EST # estimate
For help:
--help or -h print this message and exit.
--keys or -k print the list of keys and exit.
--defaults or -d print the list of keys and defaults and exit.
Other arguments are interpreted as parameter=value pairs
Notes: For GRASS 4.x, all stats are scaled up: B1 (x1000), B0 (x100),
r (x100), residuals (x100). Keep this in mind when checking
results
Running the script as r.downscale.pl -d (show defaults) results in this summary:
# #parameter =default # description (all filenames are GRASS raster maplayers) #-------------------------------------------------------------------------------- calc =BUILD # req: function performed (BUILD or EST) input =H_50 # req: BUILD & EST low res filename to be downscaled elev_lo =us_25m_dem # req: BUILD low res elev filename elev_hi =DEM30m # req: EST high res elev filename output =HIDD5d # req: EST output filename with downscaled values b1 =BONE # opt: BUILD & EST slope filename b0 =BZERO # opt: BUILD & EST intercept filename rval =RVAL # opt: BUILD corr. coeff. (r) filename resid =RESID # opt: BUILD residual (Yi - YHATi) (x10) filename ssxy =SSXY # opt: BUILD intermediate filename SSXY (sum of squares xy) ssxx =SSXX # opt: BUILD intermediate filename SSXX (sum of squares xx) ssyy =SSYY # opt: BUILD intermediate filename SSYY (sum of squares yy) d0 =1.0 # opt: BUILD wt value for 5 x 5 center cell d1 =1.0 # opt: BUILD wt value at dist. = 1 cell from center d2 =1.0 # opt: BUILD wt value at dist. = 2 cells from center e0 =8.0 # opt: EST wt value for 5 x 5 center cell e1 =2.0 # opt: EST wt value at dist. = 1 cell from center e2 =1.0 # opt: EST wt value at dist. = 2 cells from center rth =0.08 # opt: EST minimum acceptable local rsqr from build (not in orig version) #--------------------------------------------------------------------------------
For the inclusive data set, the correlation between the 4km DDs (sampled from the maplayer shown in Fig. 1 using GRASS command r.what) and actual site DDs is very good, r=0.97, reflecting the fact that DD maps were corrected using these site data. The correlation is not perfect primarily because site location precision could be improved (see lat-long coordinates). Gaussian smoothing appears to slightly improve the correlation, to r=0.998. Downscaling using the described regression algorithm also produced a high correlation, r=0.973, indicating that the method does not appear biased or inaccurate. It is expected that improvements in location precision, and in selection of a more appropriate DEM maplayer, will help with future verifications.
In the case of the exclusive data set, as expected, none of the estimated DD totals are as highly correlated with the reference (site) calculated DDs. The corrected PRISM DDs at 4km resolution show good correlation, on average within 7% of the reference data set (r=0.673). This result can be expected given the lack of precision in spatial resolution. The DDs smoothed by a gaussian filter to 500m are better correlated, r=0.764, which reflects that gaussian smoothing can be helpful for point-specific readings. The gaussian-smoothed maps do not follow the higher resolution topography, however, whereas it is known that temperature gradients will follow a much higher elevation scale of better than 100-500 m (references). The preliminary results using the downscaling method support this fact, with a correlation that is further improved to r=0.810. Much better results would be expected in a jackknife validation, by building maps using all sites other than the site tested. Such an analysis would improve the overall accuracy of these DD maps, and further support the intuitive conclusion that higher resolution maps can only be meaningfully used in conjunction with a more densely distributed network of weather stations. Also, a more rigorous test could involve weather station networks located as transects along slopes.
Table 1. Verification and validation data for 4km, gaussian smoothing, and least squares downscale techniques used for producing degree-day maps in Figs. 1-3 above. Click on station names for location map of grower-run stations.
| BUILD Model Results | ||||||||||||
| Station | Station | Station | Actual | Elev | Elev | Est DD | Est DD | Est DD | Slope | Intercept | Corr. | Residual |
| Longitude | Latitude | Name | site DD | 4km | 500m | 4km | 500m GAUSS | 500m Downscale | B1 | B0 | coeff. | Yi - Yhat |
| 1. Stations included from the analysis (verification data) | ||||||||||||
| 121:35:00.00W | 45:31:00.00N | PARO | 2647 | 600 | 528 | 2632 | 2591 | 2657 | -8.303 | 16793.34 | -.93 | -50 |
| 121:30:07.00W | 45:38:52.00N | PNGO | 2968 | 203 | 212 | 3123 | 2963 | 3189 | -11.403 | 20237.14 | -.94 | -96 |
| 121:31:01.00W | 45:41:00.00N | HOXO | 3410 | 203 | 135 | 3123 | 3315 | 3274 | -9.394 | 19639.10 | -.81 | 71 |
| 121:37:55.00W | 45:35:08.00N | DEFO | 2748 | 376 | 279 | 2751 | 2706 | 2953 | -9.060 | 18157.79 | -.86 | -62 |
| 121:12:05.00W | 45:36:01.00N | T_Dalles | 4265 | 182 | 69 | 4266 | 4335 | 4448 | -21.385 | 29003.83 | -.95 | 104 |
| correlation: EST DD vs actual DD | 0.970 | 0.998 | 0.973 | |||||||||
| 2. Hood River stations excluded from the analysis (validation data) | ||||||||||||
| 121:34:37.51W | 45:40:13.37N | Annala | 3071 | 295 | 255 | 3065 | 3138 | 3098 | -8.908 | 19211.18 | -.81 | -7 |
| 121:33:52.44W | 45:38:34.73N | Oates | 3111 | 384 | 227 | 2899 | 3025 | 3102 | -10.203 | 19209.27 | -.91 | -14 |
| 121:31:01.16W | 45:41:07.18N | MCAREC | 3257 | 203 | 155 | 3123 | 3331 | 3249 | -9.394 | 19639.10 | -.81 | 71 |
| 121:29:31.02W | 45:40:13.00N | Moore | 2837 | 203 | 208 | 3123 | 3139 | 3224 | -10.023 | 20207.59 | -.84 | 13 |
| 121:31:10.18W | 45:36:29.17N | Nakamura | 3062 | 311 | 353 | 2725 | 2708 | 2926 | -10.420 | 19214.75 | -.92 | -94 |
| 121:36:34.70W | 45:31:06.31N | Klindt | 2635 | 671 | 511 | 2490 | 2602 | 2685 | -9.261 | 17388.52 | -.92 | -34 |
| 121:34:55.54W | 45:30:39.41N | Laurance | 2422 | 600 | 534 | 2632 | 2567 | 2650 | -8.303 | 16793.34 | -.93 | -49 |
| correlation: EST DD vs actual DD | 0.673 | 0.764 | 0.810 | |||||||||
References:
Fotheringham, A. S., C. Brunsdon, and M. Charlton. 2002. Geographically Weighted Regression – the analysis of spatially varying relationships. J. Wiley & Sons.269 pp.
![]()
[Home] [Intro] [Data Table-OR] [DD Calc & Models] [Links]
![]()
On-line since April 5, 1996
This page last updated Mar 25, 2004
Contact Len Coop at coopl@bcc.orst.edu if you
have any questions about this information.